Family Trees: Generations & Propagations
Grauke attributes Psalm 90 as the ultimate reference to “generations” and the prayer focus as they pursued the SCRI grant.
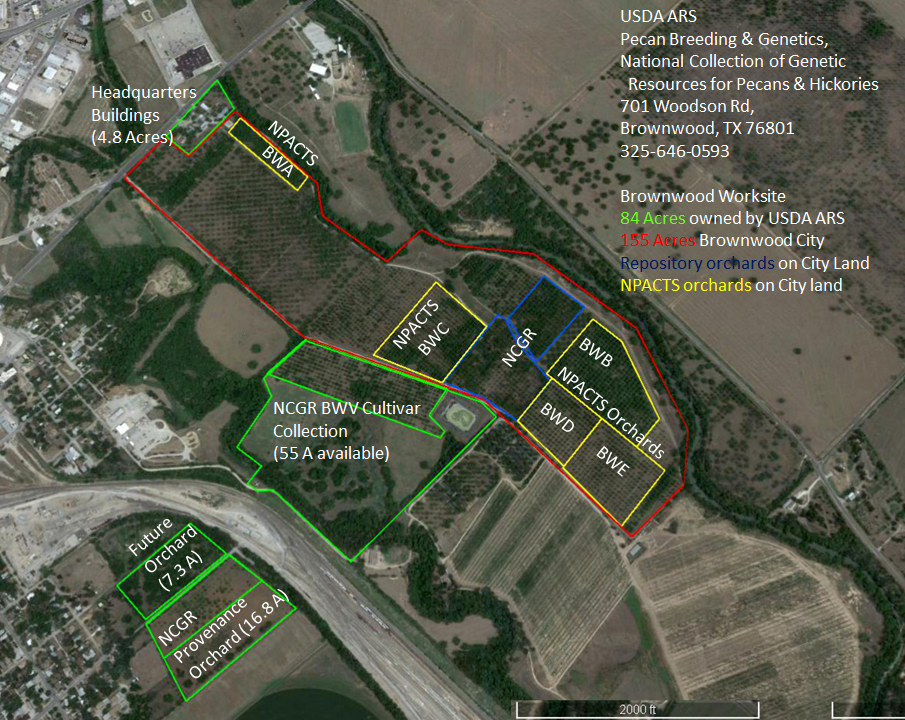
Figure 1: Brownwood worksite of the USDA ARS Pecan Breeding & Genetics Program.
Foundations of Pecan Breeding
The USDA Pecan Breeding Program began in July 1930, when appropriations were made through the efforts of Congressman James Paul Buchanan of Brenham (Romberg, 1968). Buchanan (1867-1937) was chairman of the Appropriations Committee in the U.S. House of Representatives from 1933 until his death in 1937. His seat was taken by Lyndon B. Johnson, and his name adorns Buchanan Dam and Lake Buchanan, which are well-known jewels in the Texas Hill Country Lakes. A notable pecan researcher who joined the work in that era was Louis D. Romberg (1898-1989), the first USDA pecan breeder serving from 1931 to 1968.
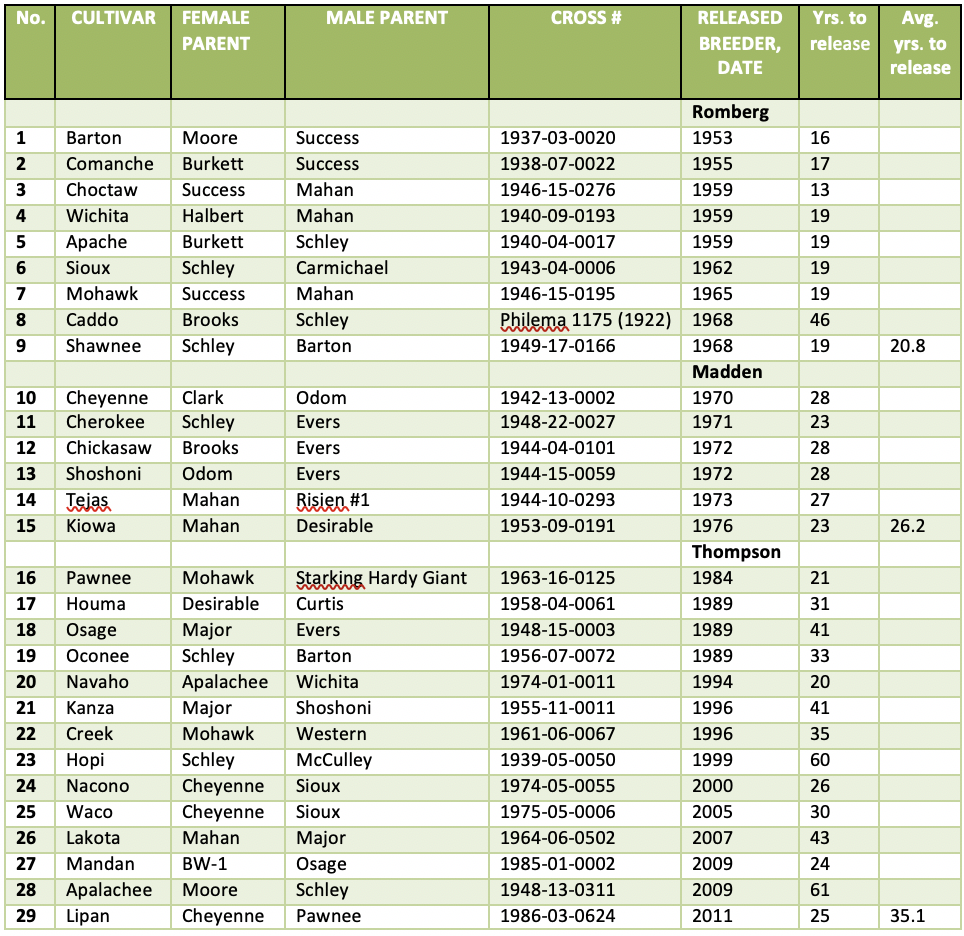
Table 1: USDA Pecan Cultivar Releases are shown with parentage, cross number, release date and years to release, and organized by breeder. (Table by Dr. LJ Grauke)
The “face” of the USDA Pecan Breeding Program has always been the cultivars it developed and released. However, the Breeding Program has suffered through very lean years: Romberg (1968) traced the early years of the program, noting a period from 1952 to 1964 when he, a technician, a part-time clerk/stenographer, and limited part-time workers were “the staff” at the Brownwood station. Despite those disadvantages, the number of years between making a cross and releasing a cultivar was lower for Romberg than subsequent breeders (Table 1).
Major changes for national pecan research were accomplished in the early 1970s when the late Ben Littlepage (1931-2018), then President of the Federated Pecan Growers Association, facilitated major funding increases to the federal budget for pecan research. Under the leadership of Howard J. Brooks, the National Program Leader for Fruit and Nut Crops for USDA Agricultural Research Service (ARS), $500,000 was redirected from other projects to pecan research, creating a budget of $1,235,000 and adding seven new scientists to the 14 ARS scientists working on pecan (Brooks, 1975). The primary locations of USDA ARS pecan research were in Byron, Georgia (now the Southeastern Fruit and Tree Nut Research Lab) and Brownwood, Texas (now a worksite of the Pecan Breeding & Genetics Research Program). The influx of capital into the USDA pecan research program allowed the hiring of two important players in that generation: Tommy E. Thompson (Research Geneticist, Texas) and Bruce W. Wood (Research Horticulturist, Georgia), both hired in 1979.
Establishment of National Fruit and Nut Repositories
Possibly the greatest change to the pecan program was the establishment of the National Fruit and Nut Germplasm Repositories (Brooks and Barton, 1977). This national plan mandated an increased scope for the program without providing substantial additional funds. The proposed plan noted that the success in breeding many crops was due to the accessibility of diverse germplasm, often “brought in from other countries since most fruit and nut crops are not indigenous to the U.S.” Individual breeding programs were often terminated by both federal and state breeding programs, with breeder’s collections being lost. The primary function of repositories was to collect, maintain, evaluate, preserve and distribute valuable genetic material, both domestically and internationally. Brooks and Barton (1977) noted that “suggested locations may not provide optimal plant growth and performance of fruit and nut clones but should provide for optimal conditions for plant survival.” (p 299). Brownwood was designated as the repository site for Carya, based on the cultivar collections L.D. Romberg had made for use as parents in his pecan breeding work (Brooks and Barton, 1977). To prepare for their new role as a National Repository, the staff at the USDA Pecan Station in Brownwood purchased 79 acres of land adjacent to the land made available to them by the City of Brownwood in a long-term Memorandum of Understanding (Figure 1).
At the first meetings of the Carya Crop Germplasm Committee (CGC) (10 July 1984), an international team composed of interdisciplinary pecan specialists from across the U.S. and Mexico made plans to collect from native pecan populations across the range of the species. Bruce Wood (USDA GA) had been trained in forestry and worked with noted forest researcher Gene Namkoong (1934-2002) to refine collection strategies and methods. Ruben Castro from Mexico’s Comisión Nacional de Fruticultura (CONAFRUT), shared information on the distribution of Mexican native pecans. Teams made nut collections across the U.S. in 1986 and across Mexico in 1987. Attempts were made to collect 100 nuts from each of five mother trees, separated by at least 200 meters. Nuts were collected from 94 trees representing 19 native populations distributed from Missouri south to Oaxaca, Mexico. Replicated orchards were planted at the Southeastern Fruit and Tree Nut Lab (SEFTNL) in Byron, Georgia with partial plantings in Burleson and Brown County, Texas, in what is now the National Collection of Genetic Resources for Pecans and Hickories (NCGR-Carya) (Grauke et al., 1989; Wood et al., 1998).
Nut collections were made relentlessly during the early 1990s to represent global populations of pecans and hickories, with some producing more valuable living collections than others. Asian species of Carya were targeted from Vietnam and China in 1990 (Grauke et al., 1991), but no living accessions represent those collections. Since few herbarium vouchers existed from Asian Carya species in U.S. herbaria, the voucher specimens collected on those trips, still maintained at the USDA Pecan Breeding and Genetics Program, were valuable for botanical characterizations (Lu et al., 1999).
Among the most valuable pecan provenance collections maintained at the College Station worksite of the USDA Pecan Breeding & Genetics worksite (Figure 2), was one grown from nuts obtained by Tommy Thompson without the effort of a plant collection. Thompson judged the 1994 Third Mexican National Pecan Show in Piedras Negras, Coahuila, Mexico and requested the nut samples that had been submitted for competition in the Native category. They were provided, along with origination and nut quality records, and were planted by maternal families in greenhouses in Brownwood. To have a representation of North American pecans in the same provenance test, nuts were collected from 16 pecan cultivars used as regional seedstocks the U.S. and planted in greenhouses. Seedlings were evaluated for growth while still in greenhouses and were planted in a randomized block test having eight blocks to capture variation in soils and to house the 1,400 seedling trees (Figure 3).
Over the next several years, tree growth and phenology were monitored seasonally and analyzed with their geographic origin. Patterns of initiation and termination of growth were related to the seedstock’s geographic origin, as were patterns of scab disease expression. Seedling diameters and heights were largest in seedlings grown from seed collected south of the orchard location but decreased to the far south and the north. Scab disease expression was greatest in southern populations, decreasing to the north. The seedling size was also reduced by the Ships clay soil in block 8, and intermediate clay soil in block 7, as compared to the Weswood silty clay loam of the first six blocks. Long-lived species like pecan will manifest the site limitations where they are grown, as well as the genetic imprint of the geographic origin of their maternal parent. Making visual observations on each of the 1,400 trees requires days, during which time growth progresses. Such test systems as this “Mexican Provenance Orchard” (CSM in Figure 2) provide an excellent laboratory for the development of improved methods of phenological monitoring using remote imaging and capturing the image and signature of the canopies at a common point in time. Such techniques would allow more frequent records of seasonal progress than possible on foot (especially without trained workers to help). Cooperative remote imaging research is currently being pursued with Sorin Popescu (TAMU).
During 1994, provenance collections were also made across Arkansas, based on the observation that no named pecan cultivars from that state were present in Repository collections (Grauke et al., 1995). Those collections were planted in a small, randomized block test, where their late seasonal growth is in stark contrast to the early growth of trees from Mexico in adjacent plantings (Figure 4).

Figure 4: Arkansas Provenance Collection (left ), Mexican Provenance Collection (right) 15 Apr. 2013.
Broadened focus, broadening tools
The development of large, randomized block tests holding diverse seedlings shifted the focus of selection from the traditional endpoint of pecan cultivar development (which was scion cultivars producing uniform, abundant, large, high-quality nuts when grafted onto seedling rootstocks) to characterizing seasonal differences in performance of seedlings over time as a function of their geographic and genetic origin. Monitoring seedling growth and development were consistent with the objectives of selecting improved seedstocks for use as rootstocks and embraced the techniques of population biologists, including foresters. Improved measures were needed to characterize the diversity being observed. Molecular markers used for population studies proved a valuable solution.
The first molecular markers developed for pecan were isozymes, multiple forms of several enzymes that catalyze important reactions. Those markers were applied to historic pecan cultivar collections (Marquard et al., 1995). Isozymes were valuable in verifying the identity of individual cultivars and were made more valuable by the knowledge of parentage provided by the breeding program’s records. Since ‘Kiowa’ was reported to be a controlled cross between ‘Mahan’ and ‘Odom’ (McCracken, 1976; Brooks and Olmo, 1978), its profile was compared with both parents, revealing that ‘Odom’ could not be the pollen parent (Marquard, 1987). With very few markers to point to the true parent, ‘Desirable’ had a profile consistent for the parent. Since not every tree is accurately labeled, confirmation of cultivar identity is significant for an individual inventory. The Pecan Breeding & Genetics program has always labeled its trees but saw the value in reporting the source trees used in molecular profiling efforts (Marquard et al., 1995, Table 2). When previously verified specimens are used for additional testing and propagation, the confidence in the results increases. Quality control in the breeding process is provided by using molecular markers. Quality control in tree labeling and at each stage of propagating is necessary for nursery producers as well. Molecular methods of cultivar verification are valuable for certification of stock tree identity. Although reliable methods of cultivar verification are now available, the certification process should be an industry-driven effort.
Isozyme analysis was also used within provenance collections maintained by NCGR-Carya (Grauke et al., 1995) and at the SEFTNL (Ruter et al. 1999). Those studies revealed genetic diversity in provenance collections not found in cultivars being used in the pecan breeding program. Genetic diversity is often recognized without understanding its function. One hint is geographic distribution. One form of the important enzyme phosphoglucose isomerase (PGI2, EC 5.3.1.9) was concentrated with native populations from West Texas (Grauke et al., 1995). Three forms, or alleles, of the enzyme are commonly found in pecan cultivars and are given the letters A, B, and C (Marquard et al., 1995). The D allele was concentrated in native populations from West Texas where it was found in 40 trees in the Lipan Creek area of Tom Green County and the Devil’s River area of Val Verde County, Texas (Grauke et al., 1995)(Figure 5a). The geographic coordinates of the mother trees from which original seed was collected are associated with living inventories in our orchards, and with isozyme profiles linked to those individual seedlings (Figure 5b). This connects accessible research orchards to the native forest and can impact developing conservation and utilization strategies.
The diversity found in wild populations of pecans and other hickories justified their inclusion in polymorphic panels, used in developing microsatellite or simple sequence repeat (SSR) markers for Carya (Grauke et al., 2003). Plastid as well as nuclear SSR markers were developed and used across provenance collections (Grauke et al., 2011) demonstrating different patterns of genetic diversity related to geographic origin revealed in the maternally inherited plastids as compared to the nuclear profile.
Across all Carya genotypes profiled by SSRs, only one individual (87MX3-2.11) grown from seed collected in Oaxaca, Mexico was found homozygous at all 14 nuclear SSR loci, possibly due to topographic isolation and inbreeding. That genotype was further evaluated by cooperators at HudsonAlpha Genome Sequencing Center and found to be “an excellent choice from which to produce a reference genome.” (Jenkins, 2015). This single individual, or 87MX2-3.11, was compared to ‘Pawnee’, which was significantly more heterozygous, and both assemblies were made available as preliminary scaffolds for continuing efforts at the 2013 ISHS Symposium where the research was reported to the pecan research community.
Coordinated Development of Genetic Tools for Pecan
The National Plant Germplasm System requires periodic analyses of challenges to crop genetic diversity from the Crop Germplasm Committees that represent them. The most recent Carya crop vulnerability report highlighted the need to cooperate domestically in developing genomic tools for use in breeding (Grauke et al., 2016). That goal was accomplished with the successful Specialty Crop Research Initiative proposal “Coordinated development of genetic tools for pecan.” This SCRI project is led by Jennifer Randall (New Mexico State University) with participation from Texas A&M University (Patricia Klein), University of Georgia (Patrick Conner), Noble Research Institute (Charles Rohla, Maria Monteros), HudsonAlpha Genome Sequencing Center (Jeremy Schmutz, Jerry Jenkins, Jane Grimwood), and USDA-ARS units working with pecans in both Georgia (Bruce Wood (retired) and Clive Bock) and Texas (LJ Grauke and Xinwang Wang). Using the template genome for sequence alignment has allowed genome-by-sequencing (GBS) analysis of pecan diversity by multiple teams. (e.g. Bentley et al., 2019). The 87MX3-2.11 template genome from the single individual mentioned above is complete with chromosome level resolution and will be published and made publicly available this year.
Teams of researchers from NMSU and TAMU working in the SCRI grant sampled 498 trees with maternal tree origins across the U.S. and Mexico in May 2019. DNA from those samples was immediately extracted and has been sent to HudsonAlpha Genome Sequencing Center for resequencing. Those genomic profiles will characterize Carya diversity in relation to geographic distribution across North America. The association of genomic markers in Carya with phenotypic traits—such as seasonal phenology, nut quality, and disease expression—linked to the individual tree records found in the Repository and Breeding Program inventories will contribute to the rapid development of marker-assisted selection.
International Cooperation
The initial HudsonAlpha draft sequence of ‘Pawnee’ was made available to Chinese cooperators in 2013. Their team completed the reference genome sequence for ‘Pawnee’, along with a reference sequence from the Chinese hickory, Carya cathayensis. Reduced-coverage genome sequence profiles were produced for many U.S. Carya species provided by NCGR-Carya (Huang et al., 2019). Comparison of gene families that differ between pecan and the Chinese hickory highlighted key genes in biotic and abiotic tolerance, oil, polyphenols, essential amino acids, and B vitamins, and provides valuable genetic perspective on the members of the genus Carya.
An important, historical botanical question relates to the relationships between species in several important plant families that are found in both eastern North America and Asia. Since the genus Carya has valuable species used as nut crops in the U.S. and China, the Chinese team was interested in exploring their genetic relationships over the extended period these species have been genetically separated. The Chinese teams were interested in exploring the diversity of all U.S. species of Carya, both diploid (with the same chromosome number as pecan) and tetraploid (with twice the number of chromosomes). All diploid hickories make fertile hybrids with pecans, but crossing tetraploids is more difficult. In their research, Wood and Grauke (2011) noted that tetraploid Carya accumulated rare earth elements. This trait was suspected to be related to adaptation to dry sites and merited further research but is currently not being pursued by any domestic researchers. Sequencing the genome of multiple tetraploids facilitates their evaluation and possibly their incorporation in breeding. For instance, Carya floridana is a tetraploid species that is well represented by collections I made across the limited geographic range of the species in Florida in 2009 (Grauke and Mendoza-Herrera, 2012). It maintains the smallest tree size of any Carya species and will be valuable to characterize as reduced tree size is pursued in breeding.
With a team of researchers lead by Jennifer Randall (NMSU) and Hiroyoshi Iwata (University of Tokyo), we sampled 1,000 accessions across all orchards at the College Station worksite in 2015. DNA from those samples was extracted by the NMSU team and analyzed using genotyping by sequencing (GBS) procedures by the Japanese team. Preliminary data is consistent with previous analyses of genetic diversity in relation to geographic distribution, highlighting the wealth of diversity available in native populations. Although the information from those analyses has been reported at scientific meetings, publications are still in preparation. It is valuable to have multiple teams, whose goals for selection are diverse, actively working cooperatively within these collections. International teams from China and Japan sought access to genetic diversity but brought their expertise, focus, and resources to the problem. The results of their work are being shared with U.S. researchers, who have contributed to the work but did not have to finance it. The research being done by the SCRI team is of the highest quality. The need to generate the funds necessary to pursue this research has delayed its delivery. One condition placed on the U.S. based research is the active involvement of the pecan industry which is achieved by an active Advisory Committee.
Conclusion
When plans for the National Fruit and Nut Repositories were being developed, pecan was an atypical crop. Unlike most fruit and nut crops grown commercially in the U.S., pecan is native to North America and is therefore surrounded by co-evolved insects and diseases. Access to native populations impacted the dynamic strategy of making provenance and species collections as well as increasing regional cultivar collections. Unlike other Repositories, our collections are actively used for breeding by the same team charged with developing and maintaining the genetically diverse collections. The close association between the USDA Pecan Breeding Program and the U.S. pecan production industry has been beneficial, providing motivation to develop methods for verifying cultivar identity that have steadily improved and developed. The goal is to develop improved cultivars to serve all diverse pecan growing regions well. Methods of population biology and forestry have been adopted and have broadened our base of diversity and increased the precision of tools available to dissect and select from that diversity. Long-term test systems have been established to control site variability that could have obscured phenotypic resolution of genetic differences. Observations have always been inventory specific, since each self-rooted seedling is unique, and each grafted inventory is either confirmed or a target for verification. The scales we use to measure diversity have benefited by the inclusion of the morphologically and genetically diverse wild relatives.
When pecan growers study Table 1, they see cultivars such as ‘Pawnee’, ‘Wichita’, ‘Oconee’ and ‘Cheyenne’ that they know and value in their orchards, despite known problems in all of them. They may not be concerned that the records of parentage for ‘Caddo’, ‘Kiowa’, and ‘Creek’ have been corrected using molecular markers. However, the excellent historical records of the breeding program, coupled with the extensive origination records of selections made from the forest by pecan growers over time, are an asset to developing markers for improved genetic selection.
When pecan growers look at the list of cultivars in Table 1 they may not notice that there are many parents, like ‘Schley’, ‘Success’, ‘Mahan’ and ‘Evers’ that occur several times, and that the offspring of those cultivars are further used as parents. Since ‘Mahan’ is a ‘Schley’ progeny, 21 of the 29 USDA releases trace back to ‘Schley.’ In more recent crosses made in the USDA Pecan Breeding Program, ‘Pawnee’ is the parent of over 20,000 progeny and ‘Wichita’ the parent of over 18,000. Although thousands of ‘Pawnee’ x ‘Wichita’ and reciprocal crosses have been evaluated in our nurseries and orchards in Texas and other states, none have been released. ‘Wichita’ by ‘Pawnee’ is the parentage of four cultivars (‘Byrd’, ‘Morrill’, ‘Cunard’ and ‘Treadwell’) patented by UGA Research Foundation (Sparks, 2010, 2013, 2014, 2015b). An additional four UGA patented cultivars (‘Huffman’, ‘Tom’, ‘Whiddon’ and ‘Tanner’) are ‘Pawnee’ by ‘Desirable’ (Sparks, 2015a, 2016, 2017, 2018).
Genetic diversity is the foundation of population survival under the challenges of the local environment, allowing for adaptation over time. Genetic diversity is the raw material used for breeding. My generation entered our work with the mandate to collect, maintain, characterize, conserve, and distribute the diversity of our crop. We have done that.
Breeding and selection target increasing uniformity. Crop wide genetic uniformity can create situations like the Southern Corn Leaf Blight Epidemic of 1969-70 that helped inaugurate the National Plant Germplasm System. It takes time to develop resistance through breeding. The temptation in commercial production is to rely on chemical protection that masks emerging problems. The U.S. pecan industry has known since 2000 that pecans get chronically infected with Pecan Bacterial Leaf Scorch (Sanderlin and Heyderich-Alger, 2000), causing yield loss (Sanderlin and Heyderich-Alger, 2003). Methods of reducing transmission of PBLS by heat treating graftwood had been reported and recommended (Sanderlin and Melanson, 2008; Melanson and Sanderlin, 2015). However, the disease is not usually noticed in well-managed orchards, and efforts to recognize or reduce it were not pursued. Sanderlin (2015) screened several pecan seedstocks and found that susceptibility to infection differed after mechanical inoculation, simulating typical insect transmission. The most susceptible seedstocks observed were the progeny of ‘Schley’, which as noted above, is the foundation parent of the U.S. pecan industry. The identification of Xylella fastidiosa in NCGR-Carya collections was precipitated by the necessity of phytosanitary certifications required for international distributions, and the inadequacy of diagnostic methods (Hilton et al., 2017; Bock et al., 2018). It was funded in part by grants from the National Plant Germplasm System. National leaders who planned the original Repository system expected multi-disciplinary teams to oversee the collections, in order to safeguard international germplasm exchange. Although Xylella fastidiosa has curtailed the international distribution of Carya graftwood from this collection, the dynamic use of these collections by international teams is facilitating cooperative use and will result in improved methods to safely share genetic diversity. There is value in recognizing disease susceptibility of both rootstocks and scion cultivars in a domestic breeding program.
The USDA ARS Pecan Breeding & Genetics Program has been involved with both the acquisition and characterization of diversity and with efforts to use it in breeding. The genetic tools that have been and are being developed within the SCRI project “Coordinated development of genetic tools for pecan” are the best path forward. Original National Plant Germplasm System plans chose sites where the target species would survive (Brooks and Barton, 1977). Trees in our collections are resilient and are surviving even though we provide them with minimal care, in order to observe their expression of disease without the filter of multiple fungicidal sprays. The developing database links the GPS coordinates of each tree to its phenotypic and genetic characterizations over time and will be valuable to foresters as well as to the U.S. improved pecan industry, and to international users who have access. These collections were envisioned as long-term assets whose dynamic use would continue. On these sites, these resilient collections with existing documentation will be here if anyone looks for them.
Cohen et al. (1991) reviewed the status of international plant germplasm conservation efforts and categorized four “eras” of off-site genetic resource conservation and use. Their timeline extended from 1850 to 2030. The period from 2010-2030 was predicted to be one of the more efficient utilization of genetic resources with closer linkage between diverse collections and breeders, facilitated by accessible, well-documented material. The National Collection of Genetic Resources for Pecans and Hickories has arrived at that point. Where the next generation goes from here will require coordination of the federal program, domestic and international research partners, and the industries who use the genus Carya. The next generation can use the genetic resources and tools developed by this generation to domesticate pecan.